Inspect model interactively#
In this notebook, we illustrate how to interactively inspect a HelicityModel using Sympy Plotting Backends and the ampform.sympy.slider module. The procedure should work for any sympy.Expr.
First, we create some HelicityModel. We could also have used pickle to load() the HelicityModel that we created in Formulate amplitude model, but the cell below allows running this notebook independently.
import qrules
from ampform import get_builder
from ampform.dynamics.builder import (
create_non_dynamic_with_ff,
create_relativistic_breit_wigner_with_ff,
)
reaction = qrules.generate_transitions(
initial_state=("J/psi(1S)", [-1, +1]),
final_state=["gamma", "pi0", "pi0"],
allowed_intermediate_particles=["f(0)(980)", "f(0)(1500)"],
allowed_interaction_types=["strong", "EM"],
formalism="canonical-helicity",
)
builder = get_builder(reaction)
builder.config.stable_final_state_ids = {0, 1, 2}
builder.config.scalar_initial_state_mass = True
initial_state_particle = reaction.initial_state[-1]
builder.dynamics.assign(initial_state_particle, create_non_dynamic_with_ff)
for name in reaction.get_intermediate_particles().names:
builder.dynamics.assign(name, create_relativistic_breit_wigner_with_ff)
model = builder.formulate()
In this case, as we saw, the overall model contains just one intensity term \(I = |\sum_i A_i|^2\), with \(\sum_i A_i\) some coherent sum of amplitudes. We can extract \(\sum_i A_i\) as follows:
import sympy as sp
amplitude = model.expression.args[0].args[0].args[0]
assert isinstance(amplitude, sp.Add)
Substitute some of the boring parameters with the provided parameter_defaults:
{\Gamma_{f_{0}(1500)},
\Gamma_{f_{0}(980)},
m_1,
m_12,
m_2,
m_{f_{0}(1500)},
m_{f_{0}(980)}}
The ampform.sympy.slider module contains some handy functions for creating ipywidgets sliders from SymPy symbols. Here is an example that will be useful when using the spb module with interactive parameter sliders.
from ampform.sympy.slider import create_slider
sliders = {
s: create_slider(s, value=v)
for s, v in model.parameter_defaults.items()
if s in amplitude.free_symbols
}
for symbol, slider in sliders.items():
if str(symbol).startswith("m"):
slider.max = 2.3
if str(symbol).startswith(R"\Gamma"):
slider.max = 1
slider.step = 0.01
display(*sliders.values())
Finally, we can can use the spb module to create a few interactive plots!
%matplotlib widget
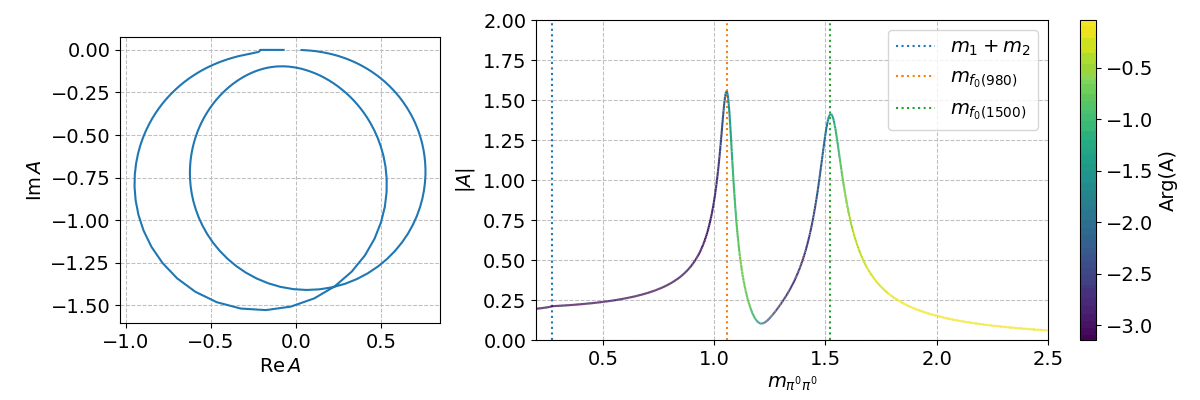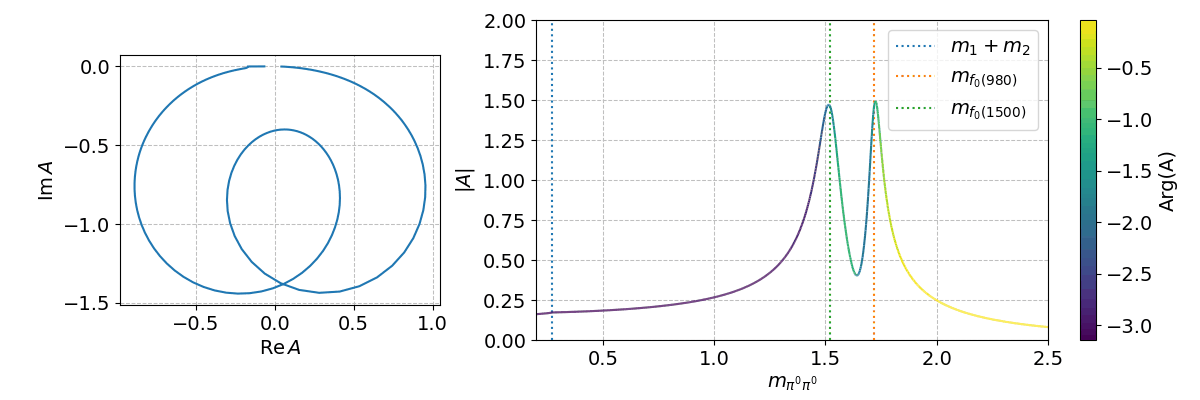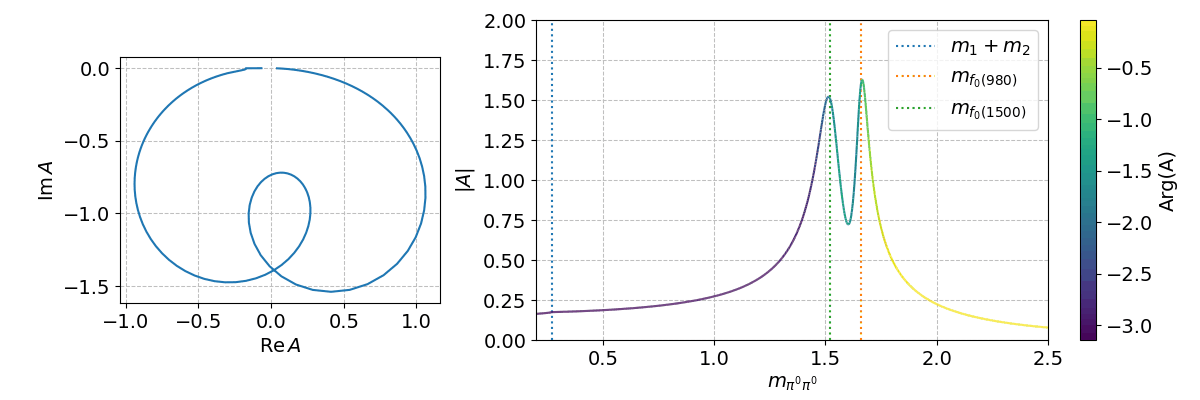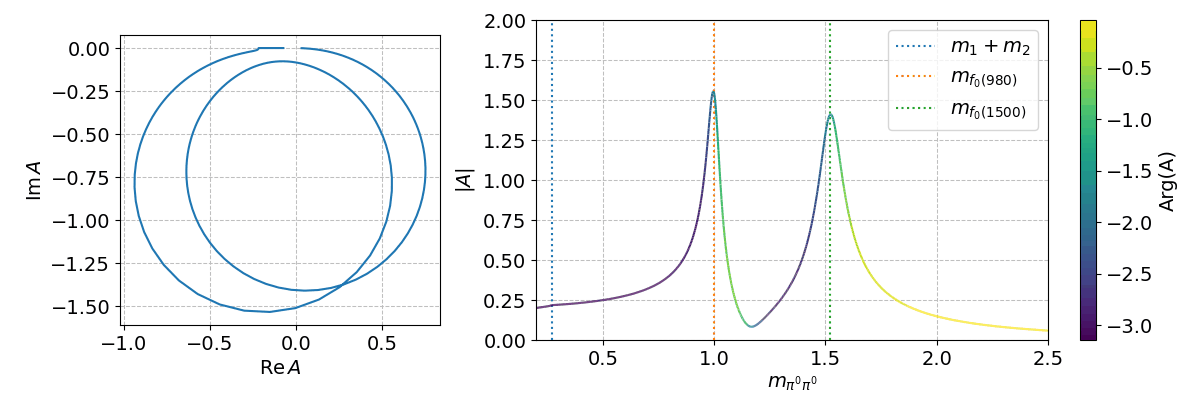
Tip
See K-matrix for why \(\boldsymbol{K}\)-matrix dynamics are better than simple Breit–Wigners when resonances are close to each other.



